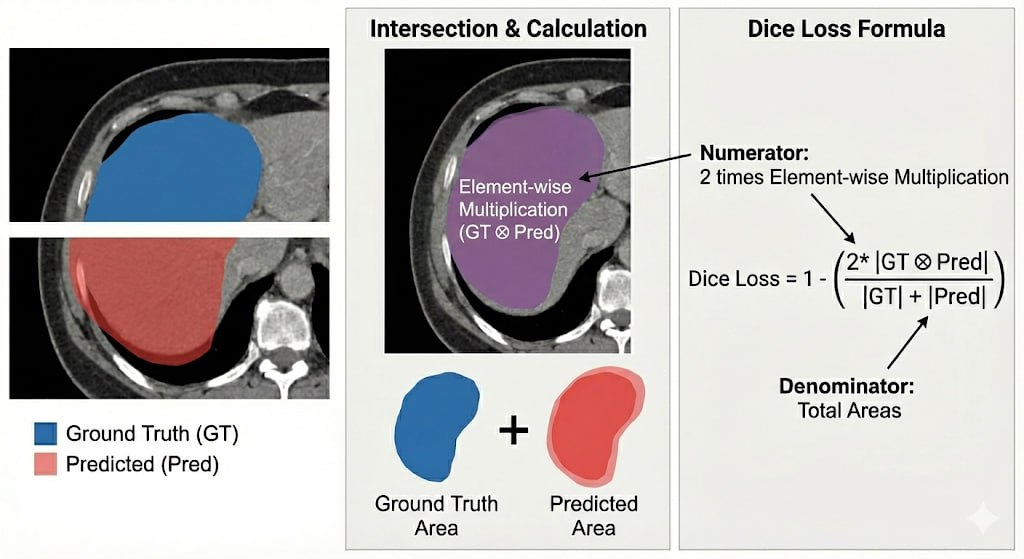
ResNetUNet vs TransUNet
We implemented and compared two state-of-the-art deep learning architectures for multi-class nuclei segmentation on the PanNuke dataset: a CNN-based ResNetUNet and a hybrid CNN-Transformer TransUNet. Both models are trained under identical conditions to segment 6 distinct cell classes in histology images from 19 tissue types.
The evaluation goes beyond simple accuracy, combining per-class Dice coefficients, paired t-tests, and False Discovery Rate (FDR) correction to rigorously assess whether observed differences between the two architectures are statistically meaningful.
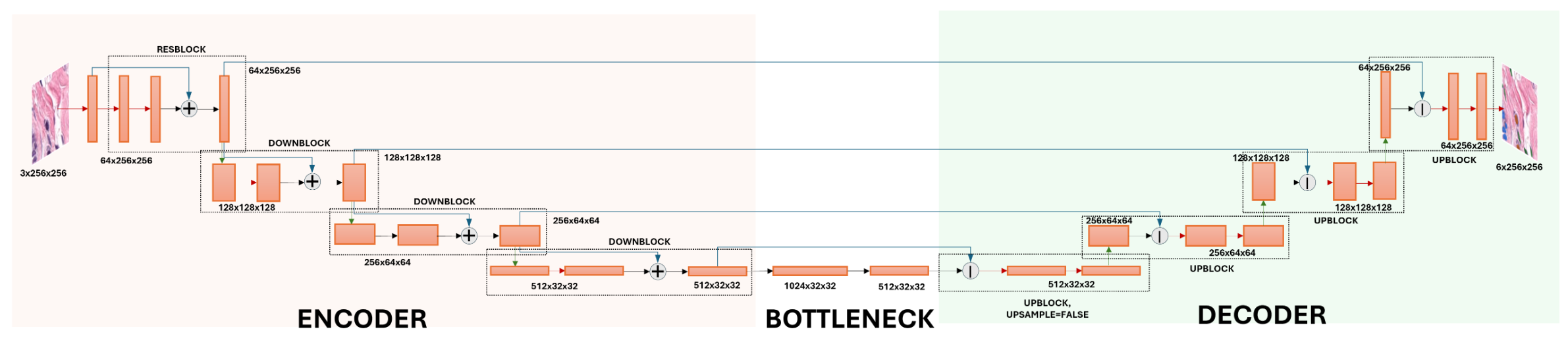
ResNetUNet Architecture
The ResNetUNet is a fully convolutional architecture combining a ResNet-inspired encoder with a progressive U-Net decoder. Skip connections fuse multi-scale features across encoder and decoder stages, enabling precise localisation of fine-grained nuclei boundaries. The model contains approximately 25M trainable parameters and uses batch normalisation and ReLU activations throughout.
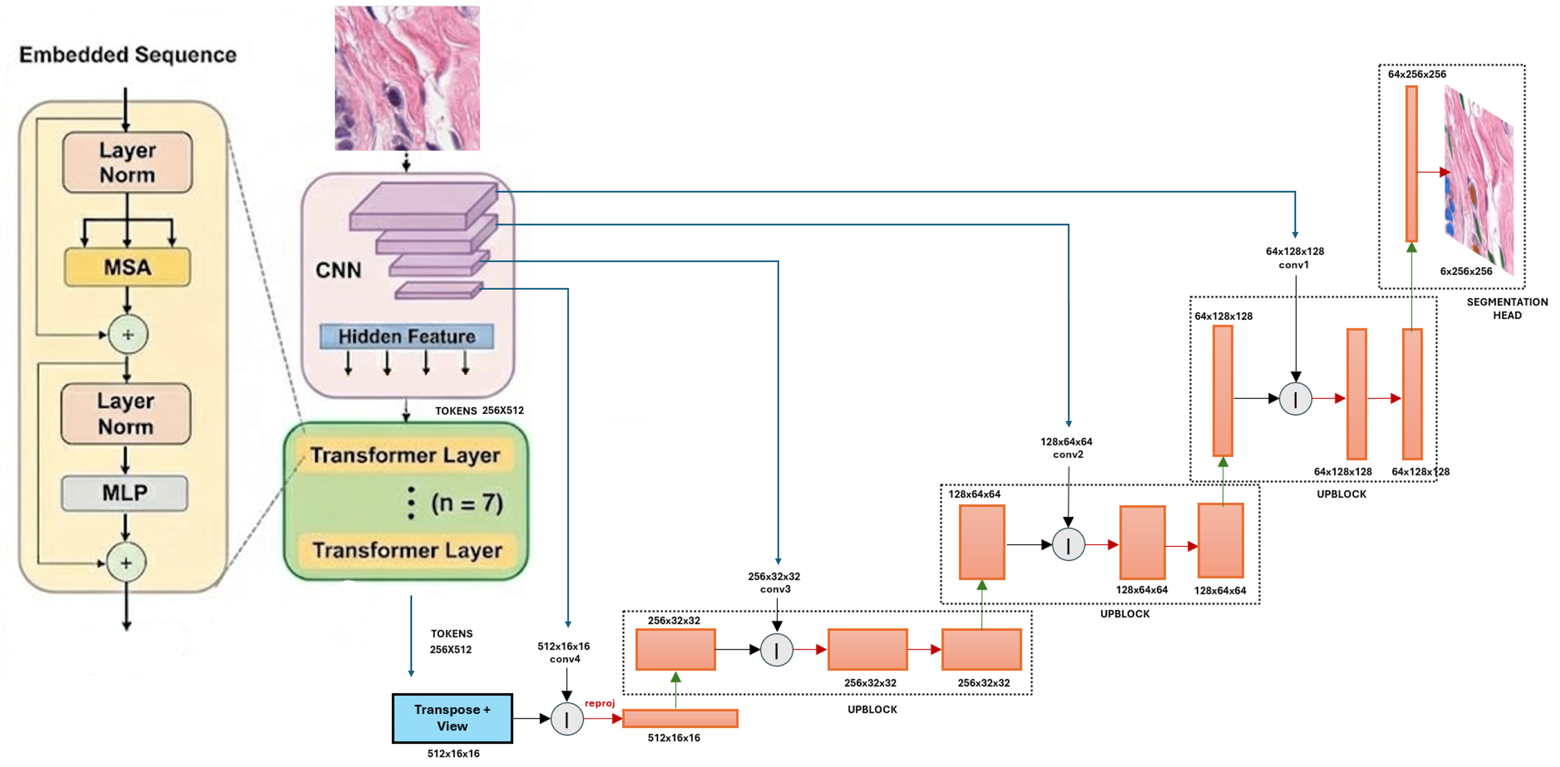
TransUNet Architecture
The TransUNet is a hybrid CNN-Transformer model that first tokenises the image into 16×16 patches via a convolutional stem, then processes them through 7 Transformer layers with 4 attention heads to capture global context. A CNN decoder with progressive upsampling reconstructs the segmentation mask. The model also contains approximately 25M trainable parameters, enabling a fair architectural comparison.
Results
TransUNet achieved a slightly higher mean Dice score (0.8410) compared to ResNetUNet (0.8390), with a statistically significant overall improvement (paired t-test: p = 0.0126). After FDR correction, class-level analysis reveals that this gain is driven primarily by Epithelial cell segmentation (+0.0162), while ResNetUNet retains a small but significant advantage on Background segmentation.